XRCC1
| View/Edit Human | View/Edit Mouse |
DNA repair protein XRCC1 also known as X-ray repair cross-complementing protein 1 is a protein that in humans is encoded by the XRCC1 gene. XRCC1 is involved in DNA repair where it complexes with DNA ligase III.
Function
| XRCC1_N | |||||||||
|---|---|---|---|---|---|---|---|---|---|
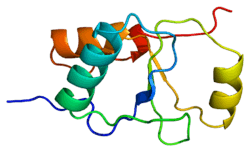
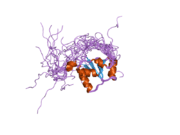
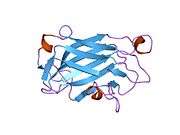 nmr solution structure of the single-strand break repair protein xrcc1-n-terminal domain | |||||||||
| Identifiers | |||||||||
| Symbol | XRCC1_N | ||||||||
| Pfam | PF01834 | ||||||||
| Pfam clan | CL0202 | ||||||||
| InterPro | IPR002706 | ||||||||
| SCOP | 1xnt | ||||||||
| SUPERFAMILY | 1xnt | ||||||||
| |||||||||
XRCC1 is involved in the efficient repair of DNA single-strand breaks formed by exposure to ionizing radiation and alkylating agents. This protein interacts with DNA ligase III, polymerase beta and poly (ADP-ribose) polymerase to participate in the base excision repair pathway. It may play a role in DNA processing during meiogenesis and recombination in germ cells. A rare microsatellite polymorphism in this gene is associated with cancer in patients of varying radiosensitivity.[3]
The XRCC1 protein does not have enzymatic activity, but acts as a scaffolding protein that interacts with multiple repair enzymes. The scaffolding allows these repair enzymes to then carry out their enzymatic steps in repairing DNA. XRCC1 is involved in single-strand break repair, base excision repair and nucleotide excision repair.[4]
As reviewed by London,[4] XRCC1 protein has three globular domains connected by two linker segments of ~150 and 120 residues. The XRCC1 N-terminal domain binds to DNA polymerase beta, the C-terminal BRCT domain interacts with DNA ligase III alpha and the central domain contains a poly(ADP-ribose) binding motif. This central domain allows recruitment of XRCC1 to polymeric ADP-ribose that forms on PARP1 after PARP1 binds to single strand breaks. The first linker contains a nuclear localization sequence and also has a region that interacts with DNA repair protein REV1, and REV1 recruits translesion polymerases. The second linker interacts with polynucleotide kinase phosphatase ( PNKP) (that processes DNA broken ends during base excision repair), aprataxin (active in single-strand DNA repair and non-homologous end joining) and a third protein designated aprataxin- and PNKP-like factor.
XRCC1 has an essential role in microhomology-mediated end joining (MMEJ) repair of double strand breaks. MMEJ is a highly error-prone DNA repair pathway that results in deletion mutations. XRCC1 is one of 6 proteins required for this pathway.[5]
XRCC1 over-expression in cancer
XRCC1 is over-expressed in non-small-cell lung carcinoma (NSCLC),[6] and at an even higher level in metastatic lymph nodes of NSCLC.[7]
XRCC1 under-expression in cancer
Deficiency in XRCC1, due to being heterozygous for a mutated XRCC1 gene coding for a truncated XRCC1 protein, suppresses tumor growth in mice.[8] Under three experimental conditions for inducing three types of cancer (colon cancer, melanoma or breast cancer), mice heterozygous for this XRCC1 mutation had substantially lower tumor volume or number than wild type mice undergoing the same carcinogenic treatments.
Comparison with other DNA repair genes in cancer
Cancers are very often deficient in expression of one or more DNA repair genes, but over-expression of a DNA repair gene is less usual in cancer. For instance, at least 36 DNA repair proteins, when mutationally defective in germ line cells, cause increased risk of cancer (hereditary cancer syndromes).[9] (Also see DNA repair-deficiency disorder.) Similarly, at least 12 DNA repair genes have frequently been found to be epigenetically repressed in one or more cancers.[9] (See also Epigenetically reduced DNA repair and cancer.) Ordinarily, deficient expression of a DNA repair enzyme results in increased un-repaired DNA damages which, through replication errors (translesion synthesis), lead to mutations and cancer. However, XRCC1 mediated MMEJ repair is directly mutagenic, so in this case, over-expression, rather than under-expression, apparently leads to cancer. Reduction of mutagenic XRCC1 mediated MMEJ repair leads to reduced progression of cancer.
Structure
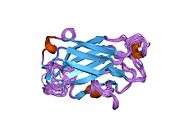
The NMR solution structure of the Xrcc1 N-terminal domain (Xrcc1 NTD) shows that the structural core is a beta-sandwich with beta-strands connected by loops, three helices and two short two-stranded beta-sheets at each connection side. The Xrcc1 NTD specifically binds single-strand break DNA (gapped and nicked) and a gapped DNA-beta-Pol complex.[10]
Interactions
XRCC1 has been shown to interact with:
References
- ↑ "Human PubMed Reference:".
- ↑ "Mouse PubMed Reference:".
- ↑ "Entrez Gene: XRCC1 X-ray repair complementing defective repair in Chinese hamster cells 1".
- 1 2 London RE (2015). "The structural basis of XRCC1-mediated DNA repair". DNA Repair (Amst.). 30: 90–103. doi:10.1016/j.dnarep.2015.02.005. PMID 25795425.
- ↑ Sharma S, Javadekar SM, Pandey M, Srivastava M, Kumari R, Raghavan SC (2015). "Homology and enzymatic requirements of microhomology-dependent alternative end joining". Cell Death Dis. 6: e1697. doi:10.1038/cddis.2015.58. PMC 4385936
 . PMID 25789972.
. PMID 25789972. - ↑ Kang CH, Jang BG, Kim DW, Chung DH, Kim YT, Jheon S, Sung SW, Kim JH (2010). "The prognostic significance of ERCC1, BRCA1, XRCC1, and betaIII-tubulin expression in patients with non-small cell lung cancer treated by platinum- and taxane-based neoadjuvant chemotherapy and surgical resection". Lung Cancer. 68 (3): 478–83. doi:10.1016/j.lungcan.2009.07.004. PMID 19683826.
- ↑ Kang CH, Jang BG, Kim DW, Chung DH, Kim YT, Jheon S, Sung SW, Kim JH (2009). "Differences in the expression profiles of excision repair crosscomplementation group 1, x-ray repair crosscomplementation group 1, and betaIII-tubulin between primary non-small cell lung cancer and metastatic lymph nodes and the significance in mid-term survival". J Thorac Oncol. 4 (11): 1307–12. doi:10.1097/JTO.0b013e3181b9f236. PMID 19745766.
- ↑ Pettan-Brewer C, Morton J, Cullen S, Enns L, Kehrli KR, Sidorova J, Goh J, Coil R, Ladiges WC (2012). "Tumor growth is suppressed in mice expressing a truncated XRCC1 protein". Am J Cancer Res. 2 (2): 168–77. PMC 3304571
 . PMID 22432057.
. PMID 22432057. - 1 2 Bernstein C, Prasad AR, Nfonsam V, Bernstein H. (2013). DNA Damage, DNA Repair and Cancer, New Research Directions in DNA Repair, Prof. Clark Chen (Ed.), ISBN 978-953-51-1114-6, InTech, http://www.intechopen.com/books/new-research-directions-in-dna-repair/dna-damage-dna-repair-and-cancer
- ↑ Marintchev A, Mullen MA, Maciejewski MW, Pan B, Gryk MR, Mullen GP (Sep 1999). "Solution structure of the single-strand break repair protein XRCC1 N-terminal domain". Nature Structural Biology. 6 (9): 884–93. doi:10.1038/12347. PMID 10467102.
- ↑ Vidal AE, Boiteux S, Hickson ID, Radicella JP (Nov 2001). "XRCC1 coordinates the initial and late stages of DNA abasic site repair through protein-protein interactions". The EMBO Journal. 20 (22): 6530–9. doi:10.1093/emboj/20.22.6530. PMC 125722
 . PMID 11707423.
. PMID 11707423. - ↑ Date H, Igarashi S, Sano Y, Takahashi T, Takahashi T, Takano H, Tsuji S, Nishizawa M, Onodera O (Dec 2004). "The FHA domain of aprataxin interacts with the C-terminal region of XRCC1". Biochemical and Biophysical Research Communications. 325 (4): 1279–85. doi:10.1016/j.bbrc.2004.10.162. PMID 15555565.
- 1 2 Gueven N, Becherel OJ, Kijas AW, Chen P, Howe O, Rudolph JH, Gatti R, Date H, Onodera O, Taucher-Scholz G, Lavin MF (May 2004). "Aprataxin, a novel protein that protects against genotoxic stress". Human Molecular Genetics. 13 (10): 1081–93. doi:10.1093/hmg/ddh122. PMID 15044383.
- ↑ Marsin S, Vidal AE, Sossou M, Ménissier-de Murcia J, Le Page F, Boiteux S, de Murcia G, Radicella JP (Nov 2003). "Role of XRCC1 in the coordination and stimulation of oxidative DNA damage repair initiated by the DNA glycosylase hOGG1". The Journal of Biological Chemistry. 278 (45): 44068–74. doi:10.1074/jbc.M306160200. PMID 12933815.
- ↑ Schreiber V, Amé JC, Dollé P, Schultz I, Rinaldi B, Fraulob V, Ménissier-de Murcia J, de Murcia G (Jun 2002). "Poly(ADP-ribose) polymerase-2 (PARP-2) is required for efficient base excision DNA repair in association with PARP-1 and XRCC1". The Journal of Biological Chemistry. 277 (25): 23028–36. doi:10.1074/jbc.M202390200. PMID 11948190.
- 1 2 Fan J, Otterlei M, Wong HK, Tomkinson AE, Wilson DM (2004). "XRCC1 co-localizes and physically interacts with PCNA". Nucleic Acids Research. 32 (7): 2193–201. doi:10.1093/nar/gkh556. PMC 407833
 . PMID 15107487.
. PMID 15107487. - ↑ Whitehouse CJ, Taylor RM, Thistlethwaite A, Zhang H, Karimi-Busheri F, Lasko DD, Weinfeld M, Caldecott KW (Jan 2001). "XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair". Cell. 104 (1): 107–17. doi:10.1016/S0092-8674(01)00195-7. PMID 11163244.
- ↑ Ewing RM, Chu P, Elisma F, Li H, Taylor P, Climie S, McBroom-Cerajewski L, Robinson MD, O'Connor L, Li M, Taylor R, Dharsee M, Ho Y, Heilbut A, Moore L, Zhang S, Ornatsky O, Bukhman YV, Ethier M, Sheng Y, Vasilescu J, Abu-Farha M, Lambert JP, Duewel HS, Stewart II, Kuehl B, Hogue K, Colwill K, Gladwish K, Muskat B, Kinach R, Adams SL, Moran MF, Morin GB, Topaloglou T, Figeys D (2007). "Large-scale mapping of human protein-protein interactions by mass spectrometry". Molecular Systems Biology. 3 (1): 89. doi:10.1038/msb4100134. PMC 1847948
 . PMID 17353931.
. PMID 17353931. - ↑ Wang L, Bhattacharyya N, Chelsea DM, Escobar PF, Banerjee S (Nov 2004). "A novel nuclear protein, MGC5306 interacts with DNA polymerase beta and has a potential role in cellular phenotype". Cancer Research. 64 (21): 7673–7. doi:10.1158/0008-5472.CAN-04-2801. PMID 15520167.
- ↑ Kubota Y, Nash RA, Klungland A, Schär P, Barnes DE, Lindahl T (Dec 1996). "Reconstitution of DNA base excision-repair with purified human proteins: interaction between DNA polymerase beta and the XRCC1 protein". The EMBO Journal. 15 (23): 6662–70. PMC 452490
 . PMID 8978692.
. PMID 8978692. - ↑ Bhattacharyya N, Banerjee S (Jul 2001). "A novel role of XRCC1 in the functions of a DNA polymerase beta variant". Biochemistry. 40 (30): 9005–13. doi:10.1021/bi0028789. PMID 11467963.
- ↑ Masson M, Niedergang C, Schreiber V, Muller S, Menissier-de Murcia J, de Murcia G (Jun 1998). "XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage". Molecular and Cellular Biology. 18 (6): 3563–71. PMC 108937
 . PMID 9584196.
. PMID 9584196.
Further reading
- Hung RJ, Hall J, Brennan P, Boffetta P (Nov 2005). "Genetic polymorphisms in the base excision repair pathway and cancer risk: a HuGE review". American Journal of Epidemiology. 162 (10): 925–42. doi:10.1093/aje/kwi318. PMID 16221808.
- Thompson LH, Brookman KW, Jones NJ, Allen SA, Carrano AV (Dec 1990). "Molecular cloning of the human XRCC1 gene, which corrects defective DNA strand break repair and sister chromatid exchange". Molecular and Cellular Biology. 10 (12): 6160–71. PMC 362891
 . PMID 2247054.
. PMID 2247054. - Thompson LH, Bachinski LL, Stallings RL, Dolf G, Weber CA, Westerveld A, Siciliano MJ (Nov 1989). "Complementation of repair gene mutations on the hemizygous chromosome 9 in CHO: a third repair gene on human chromosome 19". Genomics. 5 (4): 670–9. doi:10.1016/0888-7543(89)90107-9. PMID 2591959.
- Gyapay G, Morissette J, Vignal A, Dib C, Fizames C, Millasseau P, Marc S, Bernardi G, Lathrop M, Weissenbach J (Jun 1994). "The 1993-94 Généthon human genetic linkage map". Nature Genetics. 7 (2 Spec No): 246–339. doi:10.1038/ng0694supp-246. PMID 7545953.
- Wei Q, Xu X, Cheng L, Legerski RJ, Ali-Osman F (Nov 1995). "Simultaneous amplification of four DNA repair genes and beta-actin in human lymphocytes by multiplex reverse transcriptase-PCR". Cancer Research. 55 (21): 5025–9. PMID 7585546.
- Lamerdin JE, Montgomery MA, Stilwagen SA, Scheidecker LK, Tebbs RS, Brookman KW, Thompson LH, Carrano AV (Jan 1995). "Genomic sequence comparison of the human and mouse XRCC1 DNA repair gene regions". Genomics. 25 (2): 547–54. doi:10.1016/0888-7543(95)80056-R. PMID 7789989.
- Caldecott KW, McKeown CK, Tucker JD, Ljungquist S, Thompson LH (Jan 1994). "An interaction between the mammalian DNA repair protein XRCC1 and DNA ligase III". Molecular and Cellular Biology. 14 (1): 68–76. PMC 358357
 . PMID 8264637.
. PMID 8264637. - Trask B, Fertitta A, Christensen M, Youngblom J, Bergmann A, Copeland A, de Jong P, Mohrenweiser H, Olsen A, Carrano A (Jan 1993). "Fluorescence in situ hybridization mapping of human chromosome 19: cytogenetic band location of 540 cosmids and 70 genes or DNA markers". Genomics. 15 (1): 133–45. doi:10.1006/geno.1993.1021. PMID 8432525.
- Kubota Y, Nash RA, Klungland A, Schär P, Barnes DE, Lindahl T (Dec 1996). "Reconstitution of DNA base excision-repair with purified human proteins: interaction between DNA polymerase beta and the XRCC1 protein". The EMBO Journal. 15 (23): 6662–70. PMC 452490
 . PMID 8978692.
. PMID 8978692. - Nash RA, Caldecott KW, Barnes DE, Lindahl T (Apr 1997). "XRCC1 protein interacts with one of two distinct forms of DNA ligase III". Biochemistry. 36 (17): 5207–11. doi:10.1021/bi962281m. PMID 9136882.
- Shen MR, Jones IM, Mohrenweiser H (Feb 1998). "Nonconservative amino acid substitution variants exist at polymorphic frequency in DNA repair genes in healthy humans". Cancer Research. 58 (4): 604–8. PMID 9485007.
- Price EA, Bourne SL, Radbourne R, Lawton PA, Lamerdin J, Thompson LH, Arrand JE (Jul 1997). "Rare microsatellite polymorphisms in the DNA repair genes XRCC1, XRCC3 and XRCC5 associated with cancer in patients of varying radiosensitivity". Somatic Cell and Molecular Genetics. 23 (4): 237–47. doi:10.1007/BF02674415. PMID 9542526.
- Masson M, Niedergang C, Schreiber V, Muller S, Menissier-de Murcia J, de Murcia G (Jun 1998). "XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage". Molecular and Cellular Biology. 18 (6): 3563–71. PMC 108937
 . PMID 9584196.
. PMID 9584196. - Taylor RM, Wickstead B, Cronin S, Caldecott KW (Jul 1998). "Role of a BRCT domain in the interaction of DNA ligase III-alpha with the DNA repair protein XRCC1". Current Biology. 8 (15): 877–80. doi:10.1016/S0960-9822(07)00350-8. PMID 9705932.
- Zhou ZQ, Walter CA (Jan 1998). "Cloning and characterization of the promoter of baboon XRCC1, a gene involved in DNA strand-break repair". Somatic Cell and Molecular Genetics. 24 (1): 23–39. doi:10.1007/BF02677493. PMID 9776979.
- Taylor RM, Moore DJ, Whitehouse J, Johnson P, Caldecott KW (Jan 2000). "A cell cycle-specific requirement for the XRCC1 BRCT II domain during mammalian DNA strand break repair". Molecular and Cellular Biology. 20 (2): 735–40. doi:10.1128/MCB.20.2.735-740.2000. PMC 85188
 . PMID 10611252.
. PMID 10611252. - Marintchev A, Robertson A, Dimitriadis EK, Prasad R, Wilson SH, Mullen GP (May 2000). "Domain specific interaction in the XRCC1-DNA polymerase beta complex". Nucleic Acids Research. 28 (10): 2049–59. doi:10.1093/nar/28.10.2049. PMC 105377
 . PMID 10773072.
. PMID 10773072. - Duell EJ, Wiencke JK, Cheng TJ, Varkonyi A, Zuo ZF, Ashok TD, Mark EJ, Wain JC, Christiani DC, Kelsey KT (May 2000). "Polymorphisms in the DNA repair genes XRCC1 and ERCC2 and biomarkers of DNA damage in human blood mononuclear cells". Carcinogenesis. 21 (5): 965–71. doi:10.1093/carcin/21.5.965. PMID 10783319.
- Whitehouse CJ, Taylor RM, Thistlethwaite A, Zhang H, Karimi-Busheri F, Lasko DD, Weinfeld M, Caldecott KW (Jan 2001). "XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair". Cell. 104 (1): 107–17. doi:10.1016/S0092-8674(01)00195-7. PMID 11163244.
- Dulic A, Bates PA, Zhang X, Martin SR, Freemont PS, Lindahl T, Barnes DE (May 2001). "BRCT domain interactions in the heterodimeric DNA repair protein XRCC1-DNA ligase III". Biochemistry. 40 (20): 5906–13. doi:10.1021/bi002701e. PMID 11352725.
External links
- X-ray repair cross complementing protein 1 at the US National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the public domain Pfam and InterPro IPR002706