mir-10 microRNA precursor family
| mir-10 | |
|---|---|
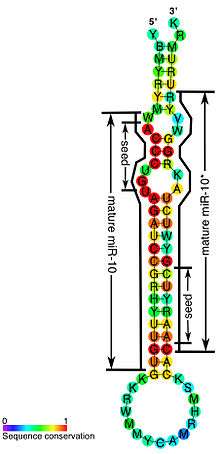 | |
| Conserved secondary structure of the mir-10 miRNA. Including marking up the mature miR and miR* sequences and corresponding seeds. | |
| Identifiers | |
| Symbol | miR-10 |
| Alt. Symbols | miR-51, miR-57, miR-99, miR-100 |
| Rfam | RF00104 |
| miRBase family | MIPF0000033 |
| HUGO | 31497 |
| OMIM | 610173 |
| Other data | |
| RNA type | microRNA |
| Domain(s) | Eukaryota; Metazoa |
The miR-10 microRNA precursor is a short non-coding RNA gene involved in gene regulation. It is part of an RNA gene family which contains miR-10, miR-51, miR-57, miR-99 and miR-100. miR-10, miR-99 and miR-100 have now been predicted or experimentally confirmed in a wide range of species.[1][2] (MIPF0000033, MIPF0000025) mir-51 and mir-57 have currently only been identified in the nematode Caenorhabditis elegans (MIPF0000268, MIPF0000271). microRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. In this case the mature sequence comes from the 5' arm of the precursor. The mature products are thought to have regulatory roles through complementarity to mRNA.
Species distribution
The presence of miR-10 has been detected in a diverse range of bilaterian animals. It is one of the most widely distributed microRNAs in animals, it has been identified in numerous species including human, dog, cat, horse, cow, guinea pig, mouse, rat, common marmoset (Callithrix jacchus), common chimpanzee (Pan troglodytes), rhesus monkey (Macaca mulatta), Sumatran orangutan (Pongo abelii), northern greater galago (Otolemur garnettii), gray short-tailed opossum (Monodelphis domestica), northern treeshrew (Tupaia belangeri), European rabbit (Oryctolagus cuniculus), African bush elephant (Loxodonta africana), nine-banded armadillo (Dasypus novemcinctus), European hedgehog (Erinaceus europaeus), lesser hedgehog tenrec (Echinops telfairi), zebra finch (Taeniopygia guttata), chicken, platypus (Ornithorhynchus anatinus), Western clawed frog (Xenopus tropicalis), Carolina anole (Anolis carolinensis), zebrafish (Danio rerio), Japanese pufferfish (Fugu rubripes), green spotted pufferfish (Tetraodon nigroviridis), Japanese killifish (Oryzias latipes), three-spined stickleback (Gasterosteus aculeatus), Florida lancelet (Branchiostoma floridae), California purple sea urchin (Strongylocentrotus purpuratus), 12 different species of fruit fly (Drosophila), Western honey bee (Apis mellifera), mosquito (Anopheles gambiae), red flour beetle (Tribolium castaneum), the nematode Caenorhabditis elegans, owl limpet (Lottia gigantea), starlet sea anemone (Nematostella vectensis) and the blood fluke Schistosoma japonicum.[3][4][5][6][7][8] In some of these species the presence of miR-10 has been shown experimentally, in others the genes encoding miR-10 have been predicted computationally.
Genomic location
The mir-10 genes are found within the Hox gene clusters. In mammals there are four Hox gene clusters, these contain five genes encoding miRNAs (mir-10a, mir-10b, mir-196a-1, mir-196a-2 and mir-196b). The mir-10a gene is located upstream of Hoxb4 and the mir-10b gene is located upstream of Hoxd4.[9] Zebrafish have seven Hox gene clusters, genes encoding miR-10 (mir-10a, mir-10b-1, mir-10b-2 and mir-10c) are found in the Hox Ba, Bb, Ca and Da clusters. A fourth miR-10 gene (mir-10d) is found elsewhere in the genome, at a location homologous to the pufferfish HoxDd cluster.[10]
miR-10*
A miRNA can be derived from each arm of the pre-miRNA hairpin. The least common of these two miRNA products is denoted by the addition of * to the miRNA name.[11] Both miR-10 and miR-10* have been detected in Drosophila. There are many potential targets for miR-10* in Drosophila, including several Hox genes, indicating that miR-10* may also be functional.[12][13] In Drosophila most mature miR-10 sequences are produced from the 3' arm of the precursor while in the beetle Tribolium castaneum most production comes from the 5' arm.[14] These changes of arm preference during evolution are termed arm switching events, and they are relatively frequent during the evolution of microRNAs.[14][15]
Pattern of expression
In adult animals, expression of miR-10 is limited to specific organs. The highest levels of miR-10a and miR-10b have been found in the kidneys of mice. Lower levels of miR-10a are seen in small intestine, lung and spleen, and lower levels of miR-10b are seen in skeletal muscle. Expression of miR-10b has also been detected in the ovaries.[7][8][16] Adult zebrafish express miR-10a in heart, testis and ovary, and miR-10b in muscle and liver.[17]
In developing embryos, miR-10 is detected at specific stages. Zebrafish embryos show miR-10a expression from 48 to 120 hours post-fertilisation, and miR-10b expression from 12 to 120 hours post-fertilisation.[17] In Drosophila expression of miR-10-3p is highest in 12- to 24-hour-old embryos and in 1st and 3rd instar larvae. Levels of miR-10-5p are highest in 12- to 24-hour-old embryos and much lower in larvae.[12]
In stage 5 Drosophila embryos (130–180 minutes post-fertilisation), miR-10 is distributed throughout 50-80% of the length of the egg. Later in development miRNA-10 becomes localised into bands, and levels decrease by stage 7 (195–200 minutes post-fertilisation). miR10 reappears by stage 11 (320–440 minutes post-fertilisation), where it is found in the ventral nerve cord, posterior midgut and hindgut. At stage 14 (620–680 hours post-fertilisation), miRNA-10 is localised to the posterior midgut and the anal pad.[18] In Drosophila larvae, miR-10-3p is found in the imaginal discs (groups of cells which are destined to become adult structures upon metamorphosis).[12] Expression of miR-10ba in mouse embryos shows a similar pattern to that of the Hoxb4 gene. Highest levels are found in the posterior trunk of the embryo, surrounding the hindlimb buds. Similarly, expression is restricted to the posterior trunk of chicken embryos.[6] In Zebrafish embryos expression of miR-10 is also restricted to the posterior trunk, later in development it is further restricted to the spinal cord.[17]
Targets of miR-10
A number of Hox genes have been shown to be regulated by miR-10. These genes encode transcription factors which are important in embryonic development. In zebrafish embryos, miR-10 binds to sites in the three prime untranslated region (3'UTR) of the HoxB1a and HoxB3a genes, which are important in anterior-posterior patterning during embryonic development. Binding of miR-10 leads to the repression of these genes. It also acts synergistically with HoxB4 to repress these genes. The mir-10 gene is located near to the HoxB1a and HoxB3a genes within the zebrafish genome, Hox-1 and Hox-3 paralogues located on different Hox clusters are not targets of miR-10.[19] Human HOXD10 gene has also been shown experimentally to be repressed by miR-10a and miR-10b.[9][20][21]
It has also been experimentally verified that miR-10a downregulates the human HOXA1 and HOXA3 genes.[21][22] Control of the Hox genes by miR-10 suggests that this microRNA may play an important role in development.[9]
In addition to the Hox genes, miR-10a represses the transcription factor USF2 and the Ran and Pbp1 genes.[23][24] The cell-surface proteoglycan Syndecan-1 is a target of miR-10b.[25]
miR-10a binds to the five prime untranslated region (5'UTR) of mRNAs encoding ribosomal proteins, and increases their translation. It binds immediately downstream of the 5' oligopyrimidine tract (5'TOP) motif, a region important in the regulation of ribosomal protein synthesis.[23]
Association with cancer
Recently there has been much interest in abnormal levels of expression of microRNAs in cancers. Upregulation of miR-10 has been found in a number of cancers. Increased levels of miR-10a have been found in glioblastoma, anaplastic astrocytomas, primary hepatocellular carcinomas and colon cancer. Increased levels of miR-10b have been found in glioblastoma, anaplastic astrocytomas, pancreatic cancer, and metastatic breast cancer.[9][20] Although high expression of miR-10b is found in metastatic breast cancers, it does not appear to be present at high levels in early breast cancers.[20][26]
Downregulation of miR-10a has been found in chronic myeloid leukemia. USF2, a target gene of miR-10a, has been found to be overexpressed in these leukemias.[24] Downregulation of miR-10a has also been found in acute myeloid leukemia, the most common acute leukemia affecting adults.[27] Conversely, miR-10a and miR-10b have found to be upregulated in acute myeloid leukemia with NPM1 mutations; these account for approximately a third of adult acute myeloid leukemia cases and contain mutations in the NPM1 gene which result in the relocation of NPM1 from the nucleus to the cytoplasm.[28] Upregulation of miR-10b has also been found in B-cell chronic lymphocytic leukemia, the most common type of leukemia.[29]
Genomic copy number abnormalities involving microRNA genes (both increases and decreases in copy number) have been found in cancers. A gain in copy number of the mir-10a gene has been found in melanoma and breast cancer.[30]
Upstream of the mir-10b gene is a promoter region containing a binding site for the Twist transcription factor (Twist). Binding of Twist to this promoter region induces miR-10b expression, leading to a reduced translation of the tumour suppressor HOXD10. This results in upregulation of RhoA/RhoC, Rho kinase activation and tumour cell invasion.[20][31]
See also
References
- ↑ Lee, R. C.; Ambros, V (2001). "An Extensive Class of Small RNAs in Caenorhabditis elegans". Science. 294 (5543): 862–4. doi:10.1126/science.1065329. PMID 11679672.
- ↑ Ambros, Victor (2001). "MicroRNAsTiny Regulators with Great Potential". Cell. 107 (7): 823–6. doi:10.1016/S0092-8674(01)00616-X. PMID 11779458.
- ↑ Li, Sung-Chou; Chan, Wen-Ching; Hu, Ling-Yueh; Lai, Chun-Hung; Hsu, Chun-Nan; Lin, Wen-Chang (2010). "Identification of homologous microRNAs in 56 animal genomes". Genomics. 96 (1): 1–9. doi:10.1016/j.ygeno.2010.03.009. PMID 20347954.
- ↑ Prochnik, Simon E.; Rokhsar, Daniel S.; Aboobaker, A. Aziz (2006). "Evidence for a microRNA expansion in the bilaterian ancestor". Development Genes and Evolution. 217 (1): 73–7. doi:10.1007/s00427-006-0116-1. PMID 17103184.
- ↑ Huang, Jian; Hao, Pei; Chen, Hui; Hu, Wei; Yan, Qing; Liu, Feng; Han, Ze-Guang; Diemert, David Joseph (2009). Diemert, David Joseph, ed. "Genome-Wide Identification of Schistosoma japonicum MicroRNAs Using a Deep-Sequencing Approach". PLoS ONE. 4 (12): e8206. doi:10.1371/journal.pone.0008206. PMC 2785426
 . PMID 19997615.
. PMID 19997615. - 1 2 Mansfield, Jennifer H; Harfe, Brian D; Nissen, Robert; Obenauer, John; Srineel, Jalagani; Chaudhuri, Aadel; Farzan-Kashani, Raphael; Zuker, Michael; Pasquinelli, Amy E; Ruvkun, Gary; Sharp, Phillip A; Tabin, Clifford J; McManus, Michael T (2004). "MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression". Nature Genetics. 36 (10): 1079–83. doi:10.1038/ng1421. PMID 15361871.
- 1 2 Teramura, M; Kobayashi, S; Hoshino, S; Oshimi, K; Mizoguchi, H (1992). "Interleukin-11 enhances human megakaryocytopoiesis in vitro". Blood. 79 (2): 327–31. PMID 1370382.
- 1 2 Beuvink, I.; Kolb, F. A.; Budach, W.; Garnier, A.; Lange, J.; Natt, F.; Dengler, U.; Hall, J.; Filipowicz, W.; Weiler, J. (2007). "A novel microarray approach reveals new tissue-specific signatures of known and predicted mammalian microRNAs". Nucleic Acids Research. 35 (7): e52. doi:10.1093/nar/gkl1118. PMC 1874652
 . PMID 17355992.
. PMID 17355992. - 1 2 3 4 Lund, A H (2009). "MiR-10 in development and cancer". Cell Death and Differentiation. 17 (2): 209–14. doi:10.1038/cdd.2009.58. PMID 19461655.
- ↑ Woltering, Joost M; Durston, Antony J (2006). "The zebrafish hoxDb cluster has been reduced to a single microRNA". Nature Genetics. 38 (6): 601–2. doi:10.1038/ng0606-601. PMID 16736008.
- ↑ "miRBase".
- 1 2 3 Ruby, J. G.; Stark, A.; Johnston, W. K.; Kellis, M.; Bartel, D. P.; Lai, E. C. (2007). "Evolution, biogenesis, expression, and target predictions of a substantially expanded set of Drosophila microRNAs". Genome Research. 17 (12): 1850–64. doi:10.1101/gr.6597907. PMC 2099593
 . PMID 17989254.
. PMID 17989254. - ↑ Stark, A.; Kheradpour, P.; Parts, L.; Brennecke, J.; Hodges, E.; Hannon, G. J.; Kellis, M. (2007). "Systematic discovery and characterization of fly microRNAs using 12 Drosophila genomes". Genome Research. 17 (12): 1865–79. doi:10.1101/gr.6593807. PMC 2099594
 . PMID 17989255.
. PMID 17989255. - 1 2 Griffiths-Jones, S.; Hui, J. H. L.; Marco, A.; Ronshaugen, M. (2011). "MicroRNA evolution by arm switching". EMBO Reports. 12 (2): 172–177. doi:10.1038/embor.2010.191. PMC 3049427
 . PMID 21212805.
. PMID 21212805. - ↑ Marco, A.; Hui, J. H. L.; Ronshaugen, M.; Griffiths-Jones, S. (2010). "Functional shifts in insect microRNA evolution". Genome Biology and Evolution. 2: 686–696. doi:10.1093/gbe/evq053. PMC 2956262
 . PMID 20817720.
. PMID 20817720. - ↑ Landgraf, Pablo; Rusu, Mirabela; Sheridan, Robert; Sewer, Alain; Iovino, Nicola; Aravin, Alexei; Pfeffer, SéBastien; Rice, Amanda; Kamphorst, Alice O.; Landthaler, Markus; Lin, Carolina; Socci, Nicholas D.; Hermida, Leandro; Fulci, Valerio; Chiaretti, Sabina; Foà, Robin; Schliwka, Julia; Fuchs, Uta; Novosel, Astrid; Müller, Roman-Ulrich; Schermer, Bernhard; Bissels, Ute; Inman, Jason; Phan, Quang; Chien, Minchen; Weir, David B.; Choksi, Ruchi; De Vita, Gabriella; Frezzetti, Daniela; Trompeter, Hans-Ingo (2007). "A Mammalian microRNA Expression Atlas Based on Small RNA Library Sequencing". Cell. 129 (7): 1401–14. doi:10.1016/j.cell.2007.04.040. PMC 2681231
 . PMID 17604727.
. PMID 17604727. - 1 2 3 Wienholds, E.; Kloosterman, W. P.; Miska, E.; Alvarez-Saavedra, E.; Berezikov, E.; De Bruijn, E.; Horvitz, H. R.; Kauppinen, S.; Plasterk, R. H. (2005). "MicroRNA Expression in Zebrafish Embryonic Development". Science. 309 (5732): 310–311. doi:10.1126/science.1114519. PMID 15919954.
- ↑ Aboobaker, A. A.; Tomancak, P; Patel, N; Rubin, GM; Lai, EC (2005). "Drosophila microRNAs exhibit diverse spatial expression patterns during embryonic development". Proceedings of the National Academy of Sciences. 102 (50): 18017–22. doi:10.1073/pnas.0508823102. PMC 1306796
 . PMID 16330759.
. PMID 16330759. - ↑ Woltering, Joost M.; Durston, Antony J.; Raible, David (2008). Raible, David, ed. "MiR-10 Represses HoxB1a and HoxB3a in Zebrafish". PLoS ONE. 3 (1): e1396. doi:10.1371/journal.pone.0001396. PMC 2148072
 . PMID 18167555.
. PMID 18167555. - 1 2 3 4 Ma, Li; Teruya-Feldstein, Julie; Weinberg, Robert A. (2007). "Tumour invasion and metastasis initiated by microRNA-10b in breast cancer". Nature. 449 (7163): 682–8. doi:10.1038/nature06174. PMID 17898713.
- 1 2 Han, L; Witmer, PD; Casey, E; Valle, D; Sukumar, S (2007). "DNA methylation regulates MicroRNA expression". Cancer biology & therapy. 6 (8): 1284–8. PMID 17660710.
- ↑ Garzon, R.; Pichiorri, F; Palumbo, T; Iuliano, R; Cimmino, A; Aqeilan, R; Volinia, S; Bhatt, D; Alder, H; Marcucci, G; Calin, GA; Liu, CG; Bloomfield, CD; Andreeff, M; Croce, CM (2006). "MicroRNA fingerprints during human megakaryocytopoiesis". Proceedings of the National Academy of Sciences. 103 (13): 5078–83. doi:10.1073/pnas.0600587103. PMC 1458797
 . PMID 16549775.
. PMID 16549775. - 1 2 Ørom, Ulf Andersson; Nielsen, Finn Cilius; Lund, Anders H. (2008). "MicroRNA-10a Binds the 5′UTR of Ribosomal Protein mRNAs and Enhances Their Translation". Molecular Cell. 30 (4): 460–71. doi:10.1016/j.molcel.2008.05.001. PMID 18498749.
- 1 2 Agirre, X.; Jiménez-Velasco, A.; San José-Enériz, E.; Garate, L.; Bandrés, E.; Cordeu, L.; Aparicio, O.; Saez, B.; Navarro, G.; Vilas-Zornoza, A.; Pérez-Roger, I.; García-Foncillas, J.; Torres, A.; Heiniger, A.; Calasanz, M. J.; Fortes, P.; Román-Gómez, J.; Prósper, F. (2008). "Down-Regulation of hsa-miR-10a in Chronic Myeloid Leukemia CD34+ Cells Increases USF2-Mediated Cell Growth". Molecular Cancer Research. 6 (12): 1830–40. doi:10.1158/1541-7786.MCR-08-0167. PMID 19074828.
- ↑ Schneider, C; Kässens, N; Greve, B; Hassan, H; Schüring, AN; Starzinski-Powitz, A; Kiesel, L; Seidler, DG; Götte, M (Nov 30, 2012). "Targeting of syndecan-1 by micro-ribonucleic acid miR-10b modulates invasiveness of endometriotic cells via dysregulation of the proteolytic milieu and interleukin-6 secretion". Fertility and Sterility. 99 (3): 871–881.e1. doi:10.1016/j.fertnstert.2012.10.051. PMID 23206733.
- ↑ Gee, Harriet E.; Camps, Carme; Buffa, Francesca M.; Colella, Stefano; Sheldon, Helen; Gleadle, Jonathan M.; Ragoussis, Jiannis; Harris, Adrian L. (2008). "MicroRNA-10b and breast cancer metastasis". Nature. 455 (7216): E8–9; author reply E9. doi:10.1038/nature07362. PMID 18948893.
- ↑ Jongen-Lavrencic, M.; Sun, S. M.; Dijkstra, M. K.; Valk, P. J. M.; Löwenberg, B. (2008). "MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia". Blood. 111 (10): 5078–85. doi:10.1182/blood-2008-01-133355. PMID 18337557.
- ↑ Garzon, R.; Garofalo, M.; Martelli, M. P.; Briesewitz, R.; Wang, L.; Fernandez-Cymering, C.; Volinia, S.; Liu, C.-G.; Schnittger, S.; Haferlach, T.; Liso, A.; Diverio, D.; Mancini, M.; Meloni, G.; Foa, R.; Martelli, M. F.; Mecucci, C.; Croce, C. M.; Falini, B. (2008). "Distinctive microRNA signature of acute myeloid leukemia bearing cytoplasmic mutated nucleophosmin". Proceedings of the National Academy of Sciences. 105 (10): 3945–50. doi:10.1073/pnas.0800135105. PMC 2268779
 . PMID 18308931.
. PMID 18308931. - ↑ Calin, G. A.; Liu, CG; Sevignani, C; Ferracin, M; Felli, N; Dumitru, CD; Shimizu, M; Cimmino, A; Zupo, S; Dono, M; Dell'Aquila, ML; Alder, H; Rassenti, L; Kipps, TJ; Bullrich, F; Negrini, M; Croce, CM (2004). "MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias". Proceedings of the National Academy of Sciences. 101 (32): 11755–60. doi:10.1073/pnas.0404432101. PMC 511048
 . PMID 15284443.
. PMID 15284443. - ↑ Zhang, L.; Huang, J; Yang, N; Greshock, J; Megraw, MS; Giannakakis, A; Liang, S; Naylor, TL; Barchetti, A; Ward, MR; Yao, G; Medina, A; o’Brien-Jenkins, A; Katsaros, D; Hatzigeorgiou, A; Gimotty, PA; Weber, BL; Coukos, G (2006). "microRNAs exhibit high frequency genomic alterations in human cancer". Proceedings of the National Academy of Sciences. 103 (24): 9136–41. doi:10.1073/pnas.0508889103. PMC 1474008
 . PMID 16754881.
. PMID 16754881. - ↑ Bourguignon, L. Y. W.; Wong, G.; Earle, C.; Krueger, K.; Spevak, C. C. (2010). "Hyaluronan-CD44 Interaction Promotes c-Src-mediated Twist Signaling, MicroRNA-10b Expression, and RhoA/RhoC Up-regulation, Leading to Rho-kinase-associated Cytoskeleton Activation and Breast Tumor Cell Invasion". Journal of Biological Chemistry. 285 (47): 36721–35. doi:10.1074/jbc.M110.162305. PMC 2978601
 . PMID 20843787.
. PMID 20843787.
Further reading
- Lagos-Quintana, M.; Rauhut, R; Lendeckel, W; Tuschl, T (2001). "Identification of Novel Genes Coding for Small Expressed RNAs". Science. 294 (5543): 853–8. doi:10.1126/science.1064921. PMID 11679670.
- Izzotti, A.; Calin, G. A.; Arrigo, P.; Steele, V. E.; Croce, C. M.; De Flora, S. (2009). "Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke". The FASEB Journal. 23 (3): 806–12. doi:10.1096/fj.08-121384. PMC 2653990
 . PMID 18952709.
. PMID 18952709. - Chen, Z; Jin, Y; Yu, D; Wang, A; Mahjabeen, I; Wang, C; Liu, X; Zhou, X (August 2012). "Down-regulation of the microRNA-99 family members in head and neck squamous cell carcinoma". Oral oncology. 48 (8): 686–91. doi:10.1016/j.oraloncology.2012.02.020. PMC 3380146
 . PMID 22425712.
. PMID 22425712.
External links
- Page for mir-10 microRNA precursor family at Rfam
- miRBase family page for mir-10
- miRBase family page for mir-99
- miRBase family page for mir-51
- miRBase family page for mir-57
- miRBase entry for human miR-10a
- miRBase entry for human miR-10b
- miRDB predicted targets for human miR-10a
- miRDB predicted targets for human miR-10b
- miRNAMap entry for human miR-10a
- miRNAMap entry for human miR-10b
- HNGC entry for miR-10a
- HGNC entry for miR-10b